[Solved] leaf disease detection using keras
-
The code given below is for image classification, it seems to be running well until the last iteration which gives the following error >
"C:/Users/admin/Desktop/plant_disease_classification/plant_disease_classification/untitled1.py", line 45, in <module>
print (label_binarizer.classes_[predict1])AttributeError: 'LabelBinarizer' object has no attribute 'classes_'
import numpy as np import pickle import cv2 from os import listdir from sklearn.preprocessing import LabelBinarizer from keras.models import load_model from keras.models import Sequential from keras.preprocessing import image from keras.preprocessing.image import img_to_array from sklearn.preprocessing import MultiLabelBinarizer from sklearn.model_selection import train_test_split import matplotlib.pyplot as plt import tensorflow default_image_size = tuple((256, 256)) #load the pickle file file_object = open(r'C:\Users\admin\Desktop\pr3\cnn_model.pkl', 'rb') model = pickle.load(file_object) #load the leaf image that is to be classified imgpath=r'C:\Users\admin\Desktop\pr1\PlantVillage\diseases\Pepper__bell___healthy\0a3f2927-4410-46a3-bfda-5f4769a5aaf8___JR_HL 8275.JPG' def convert_image_to_array(image_dir): try: image = cv2.imread(image_dir) if image is not None : image = cv2.resize(image, default_image_size) return img_to_array(image) else : return np.array([]) except Exception as e: print(f"Error : {e}") return None imar = convert_image_to_array(imgpath) np_image_list = np.array([imar], dtype=np.float16) / 225.0 label_binarizer = LabelBinarizer() predict1 = model.predict(np_image_list) print (label_binarizer.classes_[predict1])the screenshot of the above complied code and error is uploaded for reference.
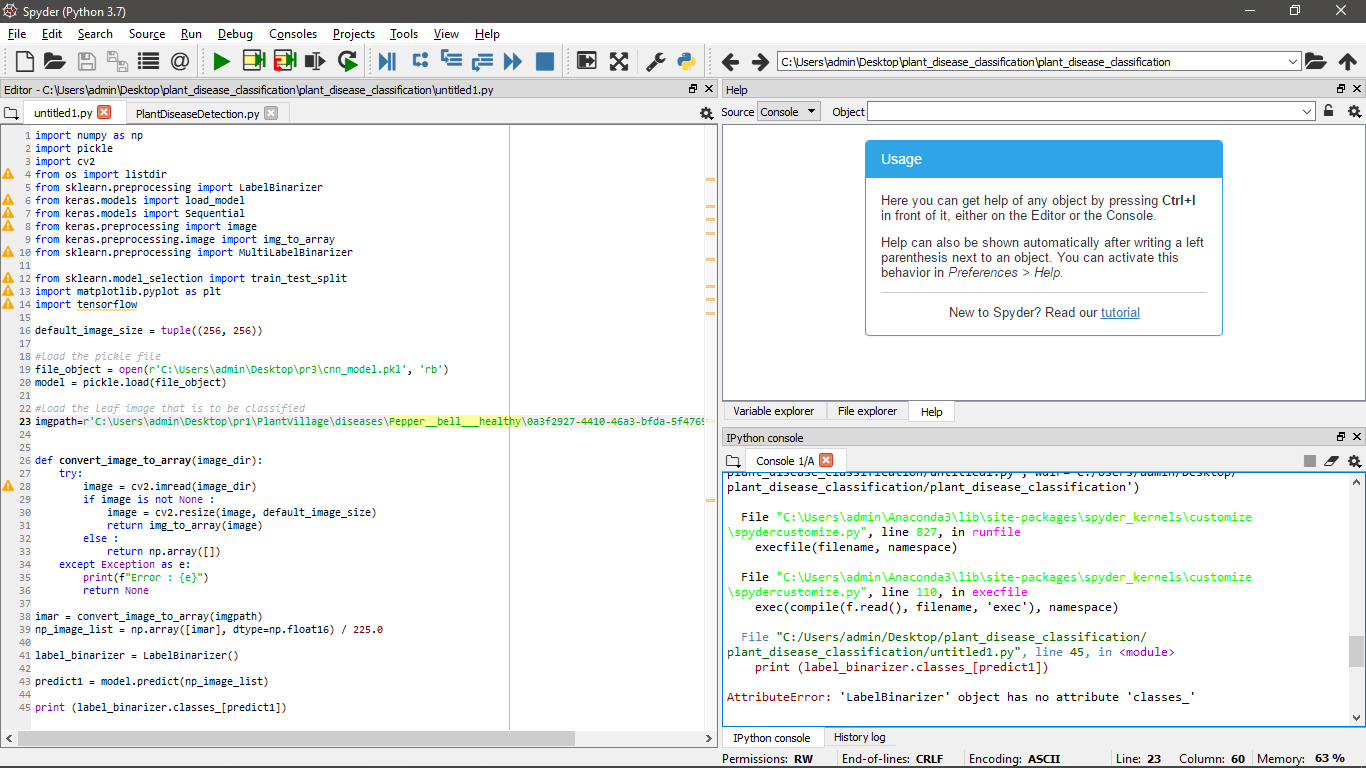
-
@sreu13 said in leaf disease detection using keras:
AttributeError: 'LabelBinarizer' object has no attribute 'classes_'
Hi, Can you try downgrade the scikit by typing
pip install scikit-learn==0.15.2and are you following any guide or something, if yes can you share that also?
-
@salmanfaris said in leaf disease detection using keras:
pip install scikit-learn==0.15.2
tried downgrading, but came up with this error.
ERROR: Failed building wheel for scikit-learn
and I've been following Kaggle kernel,
link-https://www.kaggle.com/emmarex/plant-disease-detection-using-keras -
I think you need to use the fit or fit_transform function before you predict the classes and use the Binarizer. Have a look at scikit's official documentation for the same.
https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.LabelBinarizer.html
-
i've entered this code,
def fit_transform(self, n_classes):
but as I run it, I get the indentation Error given below
File"C:/Users/admin/Desktop/plant_disease_classification/plant_disease_classification/untitled1.py", line 42
label_binarizer = LabelBinarizer()
^
IndentationError: expected an indented block -
@sreu13
fit and fit_transform are actually inbuilt functions found in the scikit-learn library. So I'd suggest you fit your model with the available data using those functions whose application can be found on scikit-learn's documentation and then proceed with the Binarizer. -
@SuperGops
so basically ,i'll have to restart and retrain the model with fit_transform? -
@sreu13 Yup
-
Could you please follow my steps:
- Uninstall your current Python 3.7 version
- Install Python 3.6.5
- If you are using spyder editor make a change to vscode.
- Go to integrated terminal of vscode and create a virtual environment
python -m venv venv- Activate your virtual environment
.\venv\Scripts\activate- Create a requirements.txt over your current working directory. Contents for requirements.txt given below:
h5py==2.8.0 imutils==0.5.1 Keras==2.2.4 Keras-Applications==1.0.6 Keras-Preprocessing==1.0.5 kiwisolver==1.0.1 matplotlib==3.0.2 numpy==1.15.3 opencv-contrib-python==3.4.3.18 Pillow==5.3.0 PyWavelets==1.0.1 scikit-image==0.14.1 scikit-learn==0.20.0 scipy==1.1.0 six==1.11.0 sklearn==0.0 tensorboard==1.12.0 tensorflow==1.12.0 termcolor==1.1.0 toolz==0.9.0- Then install necessary packages using requirements.txt file
pip install -r requirements.txt- Then run your code within this venv and say what happened as reply here.
-
@SuperGops
i also have another pickle file "label_transform", which I got as an output from referring the initial code from gaggle, is there any use of this file?label_binarizer = LabelBinarizer() image_labels = label_binarizer.fit_transform(label_list) pickle.dump(label_binarizer,open('label_transform.pkl', 'wb')) n_classes = len(label_binarizer.classes_)Kaggle >https://www.kaggle.com/emmarex/plant-disease-detection-using-keras/data


